Monday, December 8, 2025
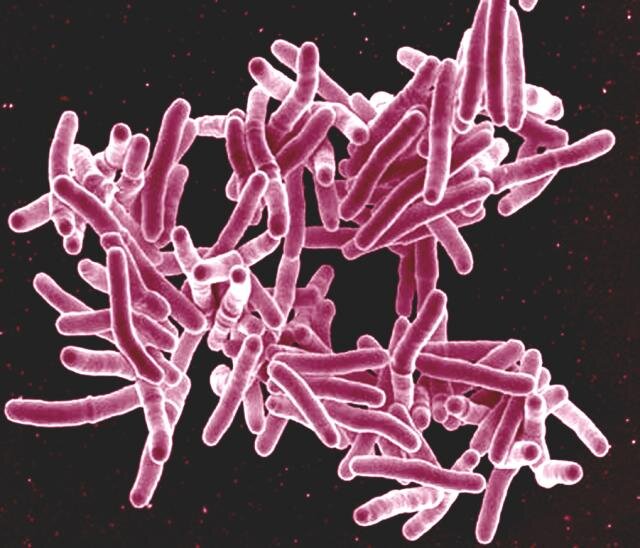
MELBOURNE — In a landmark development for the global fight against tuberculosis (TB), researchers have utilized advanced long-read DNA sequencing to reveal previously “invisible” genetic mechanisms that allow the deadly bacteria to survive antibiotic treatment.
The discovery, published this week in Nature Communications, marks a significant leap forward in genomic medicine. By constructing a comprehensive “pangenome” of Mycobacterium tuberculosis, scientists at the Peter Doherty Institute for Infection and Immunity have exposed how large structural changes in the bacterium’s DNA—often missed by standard tests—drive drug resistance. This breakthrough promises to revolutionize how clinicians diagnose and treat one of humanity’s oldest and deadliest pathogens.
The “Blind Spot” in TB Diagnostics
For the past decade, health officials have relied on “short-read” DNA sequencing to track TB outbreaks and test for drug resistance. This technology works by chopping the bacterium’s genome into tiny fragments and reassembling them like a jigsaw puzzle. While effective for detecting small, single-letter genetic mutations, this method has a critical blind spot.
“Traditional short-read sequencing struggles to accurately represent the most complex regions of the TB genome—areas full of repetitive sequences and mobile elements,” explains Aleix Canalda Baltrons, lead researcher on the study at the Doherty Institute. “Until recently, most of these large structural DNA changes were effectively invisible to us.”
These invisible changes are often driven by IS6110, a mobile genetic element or “jumping gene” that can hop around the bacterial genome. The new study reveals that IS6110 can trigger massive structural rearrangements—deletions, insertions, and inversions—that fundamentally reshape the bacterium’s biology.
Unlocking the Pangenome
Using cutting-edge nanopore “long-read” sequencing, the research team was able to read long, continuous strands of DNA, effectively bridging the gaps that baffled earlier technologies. This allowed them to move beyond the single “reference genome” traditionally used in science (based on a strain isolated over a century ago) and instead construct a pangenome—a dynamic map representing the full genetic diversity of M. tuberculosis strains from around the world.
The findings were stark:
-
Hidden Resistance: The team identified specific structural variants that delete or disable genes required for antibiotics to work. For example, if a strain deletes the gene responsible for activating a drug, the bacteria become completely immune to that treatment.
-
Regulatory Disruption: Other rearrangements were found to tweak the “volume knobs” of gene expression, allowing the bacteria to pump out toxins or shield themselves from drugs without acquiring standard mutations.
-
Evolution in Real-Time: The study showed that these structural changes occur frequently, allowing the bacteria to adapt rapidly to new antibiotic regimens.
Implications for Patient Care
The clinical implications of this discovery are profound, particularly for the estimated 400,000 patients globally who develop Multidrug-Resistant TB (MDR-TB) each year.
Currently, when a patient fails standard treatment, clinicians must embark on a months-long process of trial and error or rely on incomplete genetic data to select second-line drugs. This delay can be fatal and fuels the spread of resistant strains.
“This is the missing piece of the puzzle we’ve been looking for,” says Dr. Elena Rossi, a TB containment specialist and infectious disease physician who was not involved in the study. “We often see patients whose bacteria appear ‘susceptible’ to drugs in standard genetic tests, yet the treatment fails. This research suggests those bacteria were using these hidden structural tricks to survive. If we can see these changes upfront, we can prescribe the right drugs from day one.”
The researchers believe this technology could soon be integrated into clinical workflows. With the cost of long-read sequencing dropping rapidly, hospitals could use pangenome analysis to tailor the World Health Organization’s recommended BPaLM (bedaquiline, pretomanid, linezolid, and moxifloxacin) regimens to each patient’s specific bacterial strain.
A Persistent Global Threat
Despite being preventable and curable, tuberculosis remains a global emergency. According to the World Health Organization’s 2024 Global TB Report, the disease caused an estimated 1.25 million deaths and 10.8 million new infections in 2023 alone.
The rise of drug-resistant strains has threatened to undo decades of progress. Standard antibiotics like rifampicin and isoniazid are increasingly ineffective against these “superbug” strains, making the need for precise diagnostics more urgent than ever.
“In a disease where delayed diagnosis can have major impacts, better genomic resolution translates directly into better outcomes,” Baltrons noted. “We are moving from treating the genome as a fixed reference to viewing it as a landscape—one that changes, rearranges, and adapts.”
Challenges Ahead
While the science is promising, implementing this technology globally will face hurdles. Long-read sequencing equipment, though cheaper than before, still requires specialized laboratory infrastructure and data analysis capabilities that may be scarce in high-burden, low-resource settings.
However, the shift toward pangenomics represents a fundamental change in how we understand bacterial evolution. By shining a light on the “dark matter” of the TB genome, scientists have finally given doctors a complete map of the enemy they are fighting.
Medical Disclaimer
Medical Disclaimer: This article is for informational purposes only and should not be considered medical advice. Always consult with qualified healthcare professionals before making any health-related decisions or changes to your treatment plan. The information presented here is based on current research and expert opinions, which may evolve as new evidence emerges.
References
-
Primary Study: Canalda Baltrons, A., et al. (2025). “Genome graphs reveal the importance of structural variation in Mycobacterium tuberculosis evolution and drug resistance.” Nature Communications. [DOI: 10.1038/s41467-025-XXXXX-X]